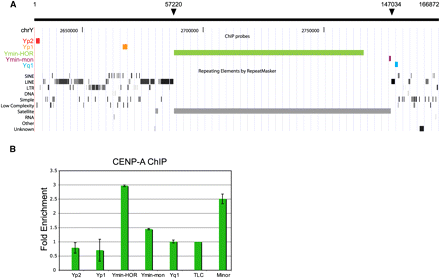
Distribution of CENPA across the Y pericentromeric domain. (A) The positions of dot blot array probes are shown against the UCSC mouse Genome Browser view (Build 37, July 2007): Yp2 and Yp1, unique DNA located on the proximal p-side of the centromere; Ymin-HOR, recognizing ∼65–79 kb of Y centromere HOR/multimeric Ymin DNA; Ymin-mon, monomeric form of Ymin at the Ymin/euchromatin boundary on the q-side of the centromere; Yq1, homologous to the Yq-arm repeat, pY353/B (Bradbury et al. 1990), and the telocentric satellite repeat, TLC. Nucleotide coordinates from BAC RP24-110P17 are annotated at the top; solid arrows mark boundaries of the Ymin array. (B) CENPA enrichment is displayed as a ratio of CENPA bound/input signal averaged over three experiments, error bars are shown as standard error. Four contiguous monomers of minor satellite DNA were used as a positive centromere DNA control (Kalitsis et al. 2003). Data are normalized against the TLC satellite repeat.











