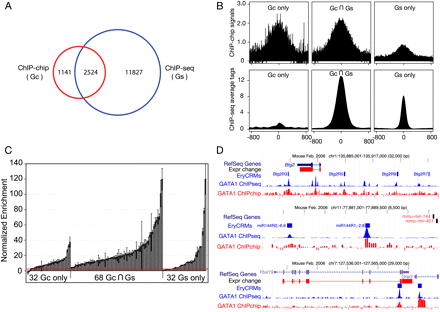
Accuracy of GATA1 peaks from high-throughput analysis of ChIPs. (A) The Venn diagram shows the relationships among peaks called from ChIP-seq and ChIP-chip data on GATA1 in G1E-ER4 cells. (Gc) G1E-ER4 ChIP-chip; (Gs) G1E-ER4 ChIP-seq. (B) Support of raw ChIP-chip and ChIP-seq data for peaks called from different technologies. The graphs show the mean ChIP-chip and ChIP-seq signals for occupancy for common and unique peaks, centered on the middle of the called peak and extending 800 bp on each side. (C) Validation of GATA1 peaks by quantitative PCR. From the peaks called for the genome-wide GATA1 ChIP data, 68 from the set common to ChIP-seq and ChIP-chip peaks, 32 from the ChIP-chip only peaks, and 32 from the ChIP-seq only peaks were chosen randomly for validation of occupancy by GATA1 using a qPCR assay, along with 20 negative control regions (not called as peaks). The bar-plot shows the mean of two determinations of the enrichment for each tested DNA segment in the GATA1 ChIP material (error bars cover the range), expressed as the number of standard deviations above the normalized mean of the negative controls (see Supplemental material). The red line indicates the threshold for validation (two standard deviations above the mean of the negative controls). (D) Data for previously studied genes, showing strong correspondence between validated erythroid CRMs and the new ChIP-seq and ChIP-chip data for GATA1. Data tracks show genes, expression response, positions of experimentally validated cis-regulatory modules, and ChIP-seq and ChIP-chip data for GATA1.











