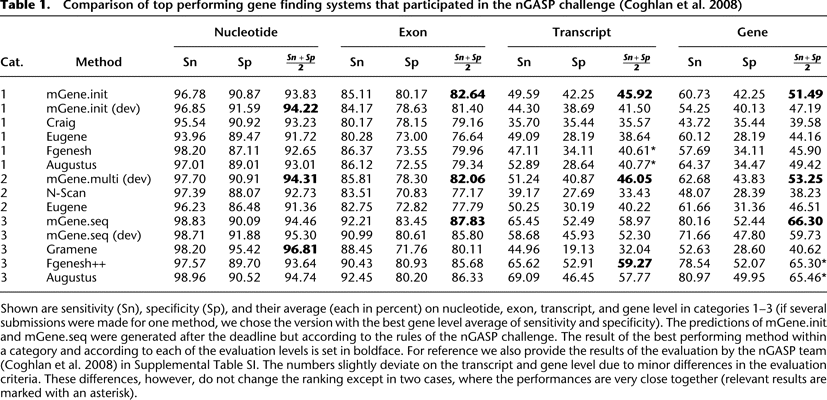
Comparison of top performing gene finding systems that participated in the nGASP challenge (Coghlan et al. 2008)
Click on table to view larger version.
-
Shown are sensitivity (Sn), specificity (Sp), and their average (each in percent) on nucleotide, exon, transcript, and gene level in categories 1–3 (if several submissions were made for one method, we chose the version with the best gene level average of sensitivity and specificity). The predictions of mGene.init and mGene.seq were generated after the deadline but according to the rules of the nGASP challenge. The result of the best performing method within a category and according to each of the evaluation levels is set in boldface. For reference we also provide the results of the evaluation by the nGASP team (Coghlan et al. 2008) in Supplemental Table SI. The numbers slightly deviate on the transcript and gene level due to minor differences in the evaluation criteria. These differences, however, do not change the ranking except in two cases, where the performances are very close together (relevant results are marked with an asterisk).











