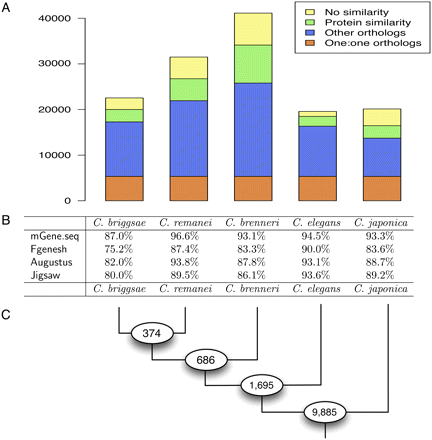
Comparison of the gene sets predicted by mGene.seq for different nematodes. (A) Number of protein-coding genes predicted for each organism and the fraction of genes with one-to-one orthologs, other orthologs with weak, and with no significant protein sequence similarity. (B) Agreement of internal exons inferred from aligned EST sequences with exons predicted by mGene.seq, Fgenesh++, Augustus, and Jigsaw. We counted a predicted exon as correct if both boundaries were correct, and as a false prediction if it overlapped a region covered by an EST alignment but did not exactly match an EST-confirmed exon. Shown is the average of sensitivity and specificity. (C) Number of orthologous groups (9885) shared among all five nematodes, as well as the number of additional orthologous groups shared across subtrees of more closely related species, which are defined by the corresponding ancestral node.











