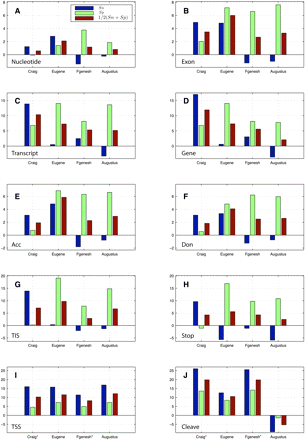
Improvement of mGene.init ab initio predictions on several evaluation levels: (A) nucleotide, (B) exon, (C) transcript, and (D) gene (each restricted to coding regions), as well as on selected signals: (E) acceptor splice sites, (F) donor splice sites, (G) TIS, (H) translation termination sites, (I) transcription start sites (TSS) (±20 nt), and (J) cleavage sites (±20 nt). mGene.init's predictions are compared with the predictions of the best submissions in category 1: Craig, Eugene, Fgenesh, and Augustus. Shown are differences of percent values for sensitivity (Sn; blue), specificity (Sp; green), and their average (red). Note that Craig and Fgenesh are not able to predict UTRs. We therefore used the predicted translation start and stop as an estimate of gene start and stop (relevant results are marked with an asterisk).











