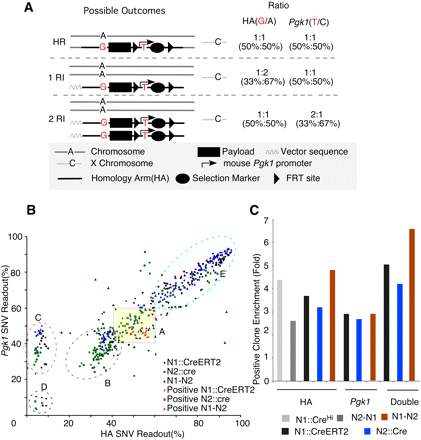
Pyroscreening of artificially introduced SNVs in targeting constructs dramatically enriched for positive HR clones. (A) Schematic illustration of pyroscreening. Artificially designed SNVs are introduced into targeting constructs: one in the HA (∼50 bp away from payload) and another one in the mouse Pgk1 promoter driving the expression of the selection marker. Different fates of the targeting construct in ES cells (HR, homologous recombination; RI, random integration) yield different ratios of the two linked SNVs to their endogenous counterparts. See text for details. (B) Pyroscreening of ES cells electroporated with (black dots) N1∷CreERT2, (green squares) N2∷cre, and (blue triangles) N1–N2. ES cells cluster into different groups. (Group A) Both HA and PGK SNV ratios fall between 40% and 60%, which suggests these are possible HR candidates. (Group B) HA SNV ratio ∼33% and PGK ∼40% suggest one copy of random integration. (Group C) The ratio suggests that these clones retained only one copy of the neomycin selection gene. (Group D) Poorly growing ES cells with heavy, G418-resistant MEF contamination. (Group E) ES cells that may have experienced multiple integrations. Note that nine out of 10 positive clones from (red dots) N1∷CreERT2, one out of one from (red square) N2∷Cre, and four of four from (red triangles) N1-N2 fall within group A. (C) When only one SNV is used for pyroscreening, the enrichment for positive clones is roughly threefold. This is increased to more than fourfold when two SNVs are combined.











