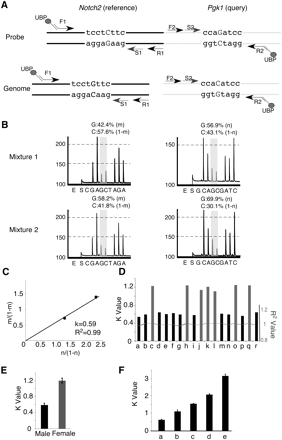
RQPS accurately determines gene copy number. (A) Illustration of the Notch2-Pgk1 probe with SNVs introduced to differentiate it from the genomic counterparts. In the RQPS probe, the reference Notch2 fragment has a G-to-C SNV while Pgk1 has a C-to-G SNV. The fragment containing the Notch2 SNV was amplified with a three-primer system: (1) the forward tailed primer F1 with a complementary region to (2) the universal biotinylated primer (UBP), and (3) the untailed reverse primer R1. The fragment containing the Pgk1 SNV was amplified with the same UBP primer and two gene-specific primers (untailed forward primer F2 and a tailed reverse primer R2). The biotinylated PCR product was purified with streptavidin-coated Sepharose beads and sequenced with a sequence primer (S1 for Notch2 and S2 for Pgk1, respectively) hybridizing near the desired SNV. (B) Example of a readout (pyrogram) from a Biotage PSQ96-MA machine. Two probe/genomic DNA mixtures were prepared in this example. Reference and query SNVs were quantified by pyrosequencing of each mixture; the relative ratio is shown. Note that a G, instead of a C, determines the ratio of the RQPS probe in each mixture because the sequencing primer for the Notch2 reference gene binds to the minus strand. (C) The plot of m/(1 − m) against n/(1 − n) gives a line that passes (0, 0) with slope k = 0.59, which suggests that there is one copy of Pgk1 in this mouse and hence it is a male. (D) RQPS accurately assigns gender by determining the copy number of the X-linked Pgk1 gene. All male mice show a k-value of ∼0.6 and all females ∼1.2. (E) Statistical analysis of k-values from 18 animals tested demonstrates the robustness of RQPS. (F) RQPS accurately measures the copy number of exon 30 from the Notch1 gene, encoding a part of the Notch1 intracellular domain, in five different lines of mice that are known to have (a) one, (b) two, (c,e) three, and (d) four copies of the NICD1, respectively. When the N1 + /▵1;Gt(ROSA)26SorNotch/Notch mouse is used, owing to a deletion removing exon 30 (query) and exon 23 (reference) from one allele (Conlon et al. 1995), three copies of N1 ICD produced k = 3.1 since a haploid reference gene was used (e).











