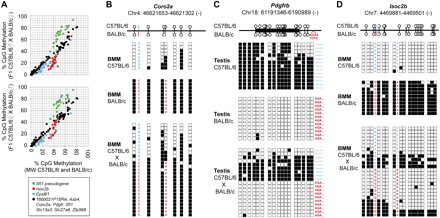
Strain-specific methylation patterns are mainly controlled in cis. (A) Averaged CpG methylation ratios of parental BMM (n = 3 for each strain) are plotted against averaged CpG methylation ratios of F1 hybrids derived from C57BL/6 (top, n = 5) or BALB/c (bottom, n = 3) sires. In eight out of 11 DMR analyzed by MALDI-TOF (marked in black), methylation patterns in F1 hybrids are almost identical to the average methylation level in parental strains (r2 > 0.97). Three DMR (Sfi1 pseudogene, Isoc2b, and Eps8l1, marked in red, green, and blue, respectively) either acquire (Sfi1 pseudogene) or lose methylation (Isoc2b, Eps8l1) in F1 hybrids relative to parental strains. (B–D) Allele-specific bisulfite sequencing of DMR in Coro2a (B, controlled in cis), Pdgfrb (C, controlled in cis), and Isoc2b (D, controlled in trans). The genomic position of CpGs within the amplicons is shown at the top. Sequence variations used to distinguish the different parental alleles are marked in blue for C57BL/6 and in red for BALB/c. Individual CpGs are represented by either white (unmethylated) or black (methylated) squares. Lines of squares represent independently sequenced clones derived from two independent sample preparations derived from reciprocal crosses. Three additional examples of DMR controlled in cis are given in Supplemental Figure S8.











