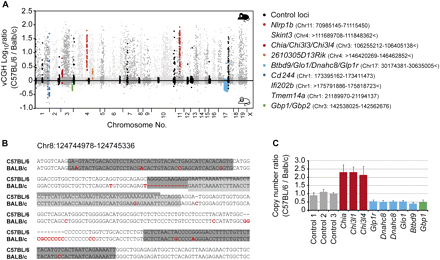
Detection of genetic variation. (A) The histogram shows a CGH-like presentation of combined signal intensities from the separate hybridizations of methylated and unmethylated DNA pools. Control loci (in black) show relatively few C57BL/6-enriched signals. Two regions (Nlrp1b and Skint, in red) are deleted in BALB/c, two regions (Chia/Chi3l3/Chi3l4 and 2610305D13Rik; in dark red and orange, respectively) are duplicated in C57BL/6, and five regions (Btbd9/Glo1/Dnahc8/Glp1r, Cd244, Ifi202b, Tmem14a, and Gbp1/Gbp2; in blue or green) appear (at least) duplicated in BALB/c. Genomic locations are provided for individual regions. The signs > and < indicate that the affected regions extended over the analyzed area. Data represent mean values of two biological replicates. (B) Sequences of an exemplary region where several probes (boxed in gray) showed C57BL/6-enriched signals. Compared to the reference strain, BALB/c contains several nucleotide exchanges (highlighted in red). (C) QPCR validation of three CNVs (Chia/Chi3l3/Chi3l4, three amplicons; Btbd9/Glo1/Dnahc8/Glp1r, five amplicons; Gbp1/Gbp2, one amplicon) detected by vCGH.











