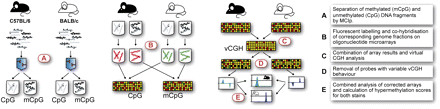
Simultaneous detection of epigenetic and genetic differences using methyl-CpG immunoprecipitation (MCIp). The experimental workflow is presented schematically. (A) Fragmented genomic DNA from bone marrow-derived macrophages of either mouse strain was fractionated using a MBD-Fc column and separated into methylated (mCpG) and unmethylated (CpG) DNA pools. (B) Both DNA pools are fluorescently labeled and compared between mouse strains by cohybridization on a locus-specific microarray using stringent conditions. (C) Array results are combined in a virtual CGH analysis to detect CNVs and sequence polymorphic regions, that are removed (D) from further analysis. (E) Differentially methylated regions are detected by analyzing remaining array probes for diametrically opposed enrichment behavior between both hybridizations (e.g., a region that is relatively enriched in the unmethylated pool of BALB/c and shows reverse enrichment behavior in the methylated pool is considered hypomethylated in BALB/c).











