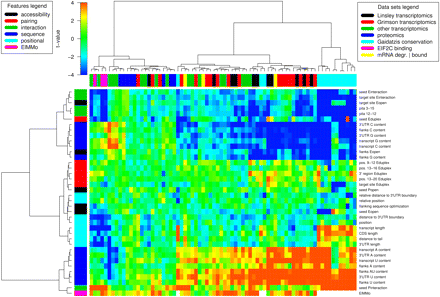
Predictive power of different features of putative miRNA target sites (rows) in predicting functional sites across the 74 data sets (columns). The data sets covered transcriptomics and proteomics measurements after miRNA transfection, transcriptomics measurements after miRNA knockdown, profiling of mRNAs bound to EIF2C/miRNA complexes, and target prediction based on comparative genomics. The heat-map shows the t-values comparing the distributions of feature values in functional vs nonfunctional miRNA target sites. The red color indicates positive predictors of miRNA functionality, while the blue color negative predictors of miRNA functionality. The dendrograms of features and data sets were produced through hierarchical clustering using Ward linkage on the Euclidean space of t-values. See also Supplemental Figure 2, in which the data set represented in each column is also indicated.











