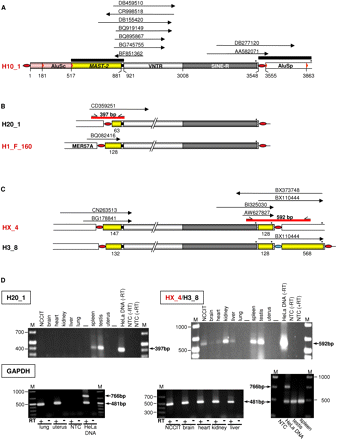
Ongoing transcription of SVA_F1 subfamily members is demonstrated by ESTs and RT-PCR analyses. (A) ESTs (arrows) aligning specifically to the source element H10_1 and its descendants (DB277120, AA582071) and to MAST2–SVA junctions of a subset of SVA_F1 subfamily members. Extensions of the queries used to identify ESTs (black bars) are shown. (B) Transcriptional control of transduction group 1 members H20_1 and H1_F_160 is effected by external promoters. Identified ESTs match 5′ junctions of the SVA elements with primary 5′ transductions. MER57A is an internal portion of the nonautonomous retrovirus 1 (ERV1). (C) ESTs indicating ongoing transcription of source element HX_4. Those ESTs that correspond to the 3′-end (BI325030, AW627827) and the 3′ junction (BX110444) of HX_4 are also consistent with transcription of the HX_4 derivative H3_8 (Fig. 7C). 3′-Ends of sense ESTs BI325030 and BX110444 as well as the 5′-end of antisense EST BX373748 are directly at the polyadenylation sites of HX_4 and/or H3_8. SVA copies highlighted in red were verified to be source elements. (D) RT-PCR analyses confirm transcription of distinct SVA_F1 subfamily members. Agarose gel electrophoresis of H20_1 and HX_4/H3_8-specific RT-PCR products derived from total RNA from human tissues and NCCIT cells. Primer binding sites and extension of the RT-PCR products are indicated as short arrows and red bars, respectively, in B and C. To control for both successful cDNA synthesis and DNA-free RNA preparations, PCRs with GAPDH-specific, intron-spanning primers (see Methods) were performed on RNA preparations from each tissue before (−RT) and after (+RT) cDNA synthesis. PCR on genomic HeLa DNA served as a positive control. The 766- and 481-bp products originate from the unspliced GAPDH gene and a processed GAPDH pseudogene, respectively (Harper et al. 2003). Sizes of obtained PCR products are indicated and correspond to expected fragment sizes. RT, reverse transcriptase; M, size marker; NTC, no template control.











