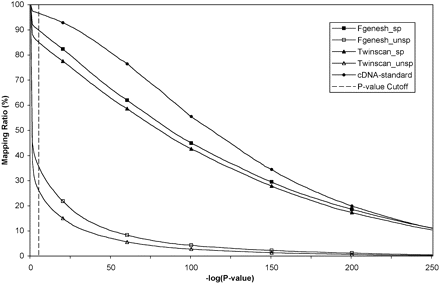
Mapping rate of rice genes on sorghum genome using TBLASTN. Rice genes predicted by Fgenesh and Twinscan are each divided into two groups: (1) supported by FLcDNA/EST (Fgenesh_sp and Twinscan_sp) and (2) not supported by FLcDNA/EST (Fgenesh_unsp and Twinscan_unsp). The genes are already filtered using MIPS TE library to remove TE-related genes. The x-axes −log(P-value) greater than 250 (P-value < 1 × 10−250) is taken as 250. The P-value cutoff 1 × 10−5 is labeled as a vertical dashed line. The confirmed genes with FLcDNA-support are included for reference purposes.











