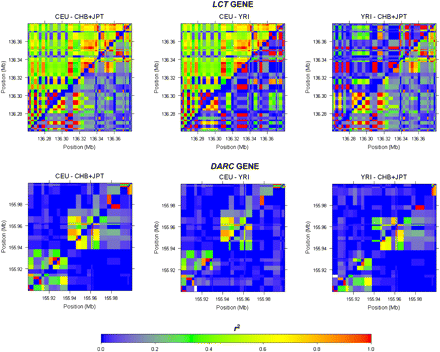
Heatmap representations of LD in two genomic regions between pairs of populations in HapMap. The upper left and lower right triangles of each plot correspond to the LD in a region for each of two populations, respectively, as measured by the pairwise r2 metric, with the plots in the first column comparing HapMap Europeans with HapMap Asians, the second column comparing HapMap Europeans with HapMap Africans, and the last column comparing HapMap Africans with HapMap Asians. The plots in the first row depict the same genomic region on chromosome 2 of 136.26 Mb–136.38 Mb spanning the LCT gene, while the plots in the second row depict the genomic region on chromosome 1 of 155.9 Mb–156.0 Mb spanning the DARC gene.











