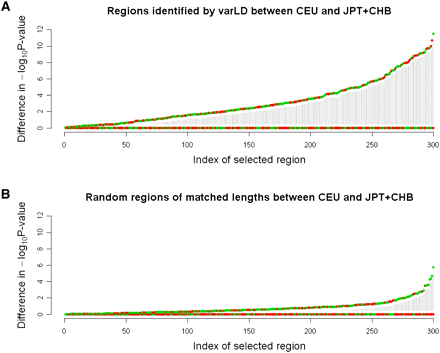
Differences in statistical evidence at the associated SNP in CEU and CHB+JPT. Comparison of the −log10P-value from a test of association between 2000 simulated cases and 2000 simulated controls at an associated SNP in each of the HapMap CEU and CHB+JPT populations. For each SNP, the larger −log10P-value is set as the baseline and is mapped to zero, and we only plot the difference of the −log10P-values. The regions are then ranked from left to right by increasing the degree of the difference in statistical evidence between CEU and CHB+JPT. (A) Three hundred randomly selected regions that have been identified by varLD to be in the top fifth percentile of the genome-wide distribution. (B) Three hundred regions that have been randomly selected across the genome, where each region spans an identical physical distance to one of the 300 varLD-identified regions from A. (Green circles) Differential statistical evidence observed in the CEU; (red circles) differential statistical evidence observed in the CHB+JPT.











