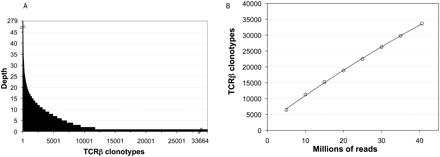
(A) TCRβ diversity. A total of 33,664 TCRβ clonotypes were identified from complete and in-frame CDR3β sequences assembled by iSSAKE. Clonotypes with a copy number of one (clonotypes identified by a single iSSAKE contig) account for 65.3% of all clonotypes. Clonotypes identified by two to 19 iSSAKE contigs account for 32.6% of all clonotypes, and high-abundance clonotypes (contig depth ≥20) account for 2.1% of the total. (B) Saturation analysis. In duplicate experiments we chose independent sets of 5, 10, 15, 20, 30, and 35 M reads at random from the pool of 40,582,229 total sequence reads in our data set. These subsets of reads were assembled and clonotypes counted as the set of complete, in-frame, nonredundant CDR3β sequences. The number of clonotypes (mean ± SD for the duplicate experiments) is plotted as a function of the number of reads. Error bars are contained within the symbols.











