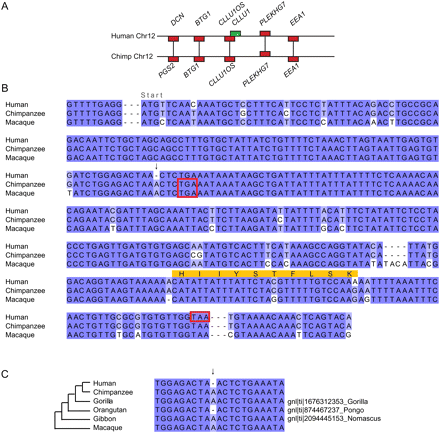
Sequence changes in the origin of CLLU1 from noncoding DNA. (A) Region of conserved synteny between human and chimp chromosomes 12. Genes are indicated by rectangular boxes and the region of chromosome is indicated by a horizontal line. Unambiguous 1:1 orthologs that were used to infer the synteny block are shown in red. One gene in this region, chronic lymphocytic leukemia up-regulated gene 1 (CLLU1), had no BLASTP hits in any other genome and is shown in green. (B) Multiple sequence alignment of the gene sequence of the human gene CLLU1 and similar nucleotide sequences from the syntenic location in chimp and macaque. The start codon is located immediately following the first alignment gap, which was inserted for clarity. Stop codons are indicated by red boxes. The sequenced peptide identified from this locus is indicated in orange. The critical mutation that allows the production of a protein is the deletion of an A nucleotide, which is present in both chimp and macaque (indicated by an arrow). This causes a frameshift in human that results in a much longer ORF capable of producing a 121-amino acids-long protein. Both the chimp and macaque sequences have a stop codon after only 42 potential codons. (C) Alignment of the region around the critical human enabler-mutation with similar nucleotide sequences from the syntenic regions in chimp, and macaque and sequence traces from gorilla, gibbon, and orangutan. For gorilla, gibbon, and orangutan the trace database accession number is shown on the right. The disabler is also shared by gorilla and gibbon indicating it is ancestral.











