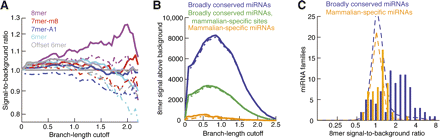
Conservation of sites matching mammalian-specific miRNAs. (A) Signal-to-background ratio for sites matching 53 mammalian-specific miRNA families in 18 placental mammals, otherwise as in Figure 2D. (B) Signal above background for 8mer sites matching either broadly conserved or mammalian-specific miRNAs in 18 placental mammals, otherwise as in Figure 2F. Analysis of 8mer sites matching broadly conserved miRNAs considers either all sites (blue) or excludes those sites conserved beyond placental mammals (green). (C) Signal-to-background ratios for 8mer sites matching individual miRNAs in orthologous 3′UTRs of placental mammals at optimal sensitivity (branch-length cutoff of 0.85). For the broadly conserved miRNA set, conservation signal excludes sites conserved beyond placental mammals. Distributions expected if miRNA targeting conferred no preferential conservation were estimated using the average signal-to-background ratio of 8mer controls selected for each site, considering GC content and dinucleotide-based conservation (broken lines). Expectations differed between the two sets because of different miRNA numbers and different dinucleotide compositions.











