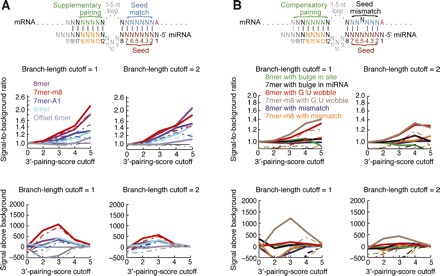
Conserved pairing to the 3′ ends of miRNAs. (A) Preferential occurrence of pairing to the 3′ region of the miRNA, which can supplement canonical sites (diagrammed at top). Signal-to-background ratio (top two graphs) and signal above background (bottom two graphs) are plotted at conservation branch-length cutoffs of 1.0 (left) and 2.0 (right). Five percent confidence limits are shown as dashed lines. (B) Preferential occurrence of pairing to the 3′ region of the miRNA, which can compensate for mismatches or bulges in seed pairing (diagrammed at top). As in A, except that seed matches were replaced as indicated with sites containing a mismatched, G:U wobble, or bulged nucleotide.











