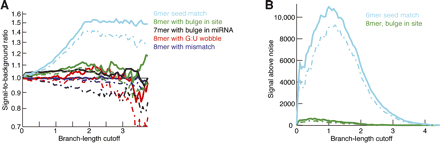
Occasional preferential conservation of imperfect sites. (A) Signal-to-background ratio for sites with the indicated single-nucleotide mismatches and bulges. A mismatch or G:U wobble must occur opposite miRNA seed nucleotides 2–7. A bulge in the site must occur between bases that pair to consecutive seed nucleotides 2–7, and a site creating a bulge must involve a 7mer match that skips one of the seed nucleotides 2–7. Results for the canonical 6mer site (Fig. 2D) are included for comparison. Broken lines indicate 5% lower confidence limit (z-test). (B) Weak signal above background for a class of imperfect sites found to have a significantly positive signal-to-background ratio. Results for the canonical 6mer site (Fig. 2F) are included for comparison. Broken lines indicate 5% lower confidence limit (z-test).











