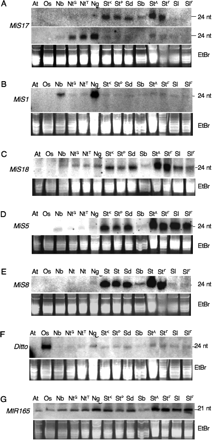
MiS small RNAs detected in Solanum and Nicotiana species by Northern blot hybridization. Forty micrograms of small RNA-enriched from total RNA isolated from Solanaceae species was size separated using a 20% acrylamide gel, blotted, and hybridized with MiS-derived probes (see Supplemental Table S3.1 for a description of probes). (A) Hybridization probe for (upper panel) Solanum-MiS17 probe; (lower panel) Nicotiana-MiS17; (B) Nicotiana-MiS1 probe; (C) Solanum-MiS15 probe; (D) Solanum-MiS5 probe; (E) Solanum-MiS8 probe; (F) Os-Ditto probe; (G) MIR165. An EtBr stain is shown for each gene as a sample loading control. (At) A. thaliana, Col-0; (Os) O. sativa Nipponbare; (Nb) N. benthamiana; (NtS) N. tabacum, SR1; (NtTG34) N. tabacum, SR1:NN; (Ng) N. glutinosa; (StKE) S. tuberosum, Kennebec; (StPI) S. tuberosum, Pentland Ivory; (Sd) S. demissum; (Sb) S. bulbocastanum; (StL) S. tuberosum, Katahdin leaves; (StF) S. tuberosum, Katahdin flowers; (SlL) S. lycopersicum VF36 leaves; (SlF) S. lycopersicum, VF36 flowers.











