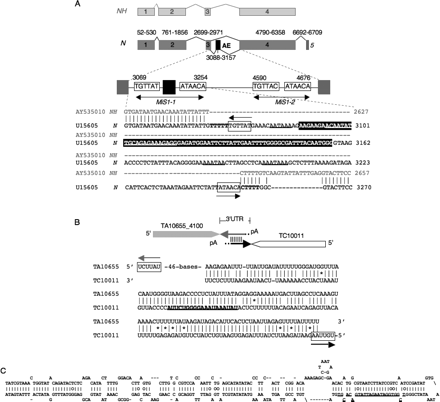
MiS1 is expressed as an alternative exon and as a 3′-UTR. (A) The structure of the N gene (U15605) and a candidate N ortholog, NH (A506320), showing (boxes) exons and (angled lines) introns. The full-length Ns transcript includes (gray boxes) exons 1–5. The alternatively spliced NL transcript includes exons 1–5 and the (AE; black box) alternative exon and is predicted to encode a truncated product by an AE-encoded frameshift. Intron 3 has two MITE insertions (MiS1-1 and MiS1-2). The entire AE is located within MiS1-1. The insertion of MiS1-1 in N is shown relative to the RESite in NH; with the AE highlighted in black. (Boxes) MiS1 TIRs; (numbers) indicate nucleotide positions in base pairs. (Bold, underlined lettering) A sequence complementary to 21 nt of MiS1 N. glutinosa siRNA, sNg_317-22-1 (5′-TCCTCTTTCTCTGCAATATTGT-3′), located within the alternative exon of MiS1-1. (Underline) Putative polyadenylation termination sequences. (B) MiS1 elements inserted in opposite orientations in different genes. (Boxes with arrowheads) The coding regions of two N. benthamiana transcript assemblies, TA10655_4100 (http://plantta.tigr.org/cgi-bin/plantta_report.pl?ta=TA10655_4100) and TC10011 (http://compbio.dfci.harvard.edu/tgi/cgi-bin/tgi/tc_report.pl?tc=TC10011&species=n_benthamiana). They are predicted to encode a hypothetical protein and WRKY transcription factor 22, respectively (based on BLASTX searches of GenBank) (see Supplemental Table S2.1). (Gray and black lines with arrowheads showing orientation) A MiS1 element is inserted in the 3′-UTR of each transcript in complementary orientations. The sequences of the MiS1 insertions are shown, indicating the putative dsRNA formed between TA_10655_4100 and TC10011. (Boxes) MiS1 TIRs. Forty-six bases of TA10655_4100 are omitted. (Bold, underlined lettering in TC10011) A sequence complementary to N. glutinosa MiS1 siRNA, sNg_1919_19_1 (5′-TAAGACCCCTTTATTTATA-3′) (Table 2). (C) A predicted (MFOLD) hairpin structure of the reverse complement of the MiS9 sequence within S. tuberosum EST DN938187 (405–108 nt) is shown (Supplemental Table S2.1). DN93187 shares homology with Solanum ESTs whose predicted products exhibit strong similarity to the C-terminal portion of Arabidopsis DCL2 (Supplemental Table S2.1). (Bold, underlined lettering in DN93187) A sequence complementary to S. demissum MiS9 siRNA, sSd_2627_24_1 (5′-TAAGACCCCTTTATTTATA-3′) (Table 2).











