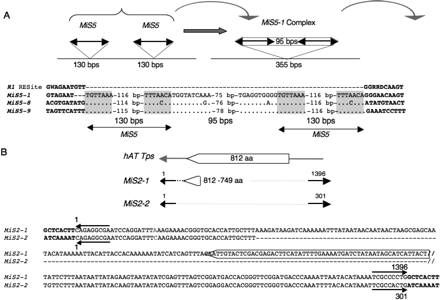
Structures of MiS5 and MiS2. (A) Several MiS5 members are found as complex with an intervening sequence. A model is presented in which tandem insertions of the same MiS5 element generate a MiS complex that captures and transposes intervening sequence. (Bold black lines flanked by arrowheads, TIRs) MiS. The sequences of the MiS5-1 RESite in S. demissum (R1; AC149265.1) and three MiS5 complexes, MiS5-1 in S. demissum (AC149266, 1364–1718), MiS5-8 in S. lycopersicum (AF411809, 126,063–126,418), and MiS5-9 in S. tuberosum (TA45582_4113, 250–606), are shown. (Dots) Nucleotide identity; (dashes in R1) the RESite lacking MiS5 insertion; (dashes in MiS5-1) a 3-bp deletion at its insertion site; (gray) TIRs; (bold) flanking sequence. (B) MiS2-1 (1396 bp) and MiS2-2 (301 bp) are hAT-type elements in GenBank accessions AC149265, 50,964–52,375, and AC149488, 68,757–69,073, respectively. (Open arrowhead) The MiS2-1 sequence is predicted to encode a truncated product with similarity to the C terminus (749–812 amino acids) of candidate S. demissum hAT transposase (AAT39314). Conserved sequences between MiS2-1 and MiS2-2 are shown aligned. (Horizontal arrows above and below the aligned sequences) The 9-bp MiS2 TIRs. (Bold) The 8-bp TSDs generated by the MiS2 insertions. The MiS2-1 sequence enclosed in the open arrow is predicted to encode amino acids 794–812 of the candidate S. demissum hAT transposase.











