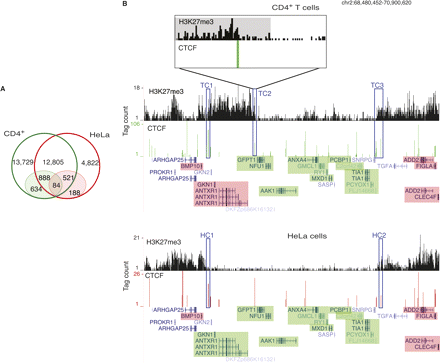
Barrier CTCF sites are cell-type-specific. (A) While over 50% and 74% of the CTCF-binding sites in CD4+ T cells and HeLa cells, respectively, overlap (open circles), only 5% and 11% of the barrier CTCF sites in the respective cell types overlap. The barrier CTCF sites are shaded. (B) A 2.5-Mbp region in chromosome 2 in CD4+ T cells (top) and HeLa cells (bottom). (Green) Expressed genes; (pink) silent genes. (HC1, HC2) Barrier CTCF sites in HeLa cells. A locus, which is occupied by CTCF in both CD4+ T cells and HeLa cells, could be functioning as a barrier in HeLa cells (HC1), but not in CD4+ T cells (TC1). (TC2, TC3) CTCF-binding sites, which could potentially be playing the role of domain barriers in CD4+ T cells. (Inset) An enlargement of the 30-kb region (TC2) in CD4+ T cells. The H3K27me3 domain is shaded gray.











