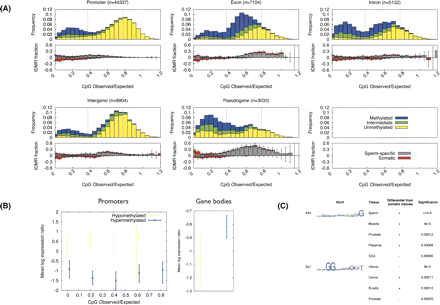
Analysis of tissue-specific differentially methylated regions (tDMRs). (A) The top half of each panel shows the DNA methylation profiles observed in sperm. In the bottom half of each panel, bars above the line represent the proportion of ROIs in each CpGo/e category that display <40% methylation in sperm, but >60% methylation in all somatic tissues (gray) or >60% methylation in one or more (but not all) somatic tissues (red, i.e., these are somatic tDMRs). Bars below the line (i.e., negative values) represent the proportion of ROIs in each CpGo/e category that display >60% methylation in sperm, but <40% methylation in all somatic tissues (gray) or <40% methylation in one or more (but not all) somatic tissues (red, i.e., these are also somatic tDMRs). (B) Comparison of tissue-specific DNA methylation and gene expression. (Left) Comparison of promoter DNA methylation (only ROIs overlapping the TSS were used) and gene expresson between whole blood and uterus. Gene expression data are from GNF SymAtlas database (Su et al. 2004). Insufficient data were available for CpGo/e > 0.8. Yellow bars represent genomic regions that display <40% methylation in whole blood and >60% methylation in uterus. Blue bars represent genomic regions that display >60% methylation in whole blood and <40% methylation in uterus. (Right) Comparison of gene-body methylation with gene expression of the associated gene. Intronic and exonic methylation data were combined for this analysis. All ROIs in these categories were used, not just CpG-dense ROIs, but we did not stratify by CpGo/e since there were insufficient numbers of exonic/intronic CpG-poor ROIs. Both figures show 95% confidence interval. The color code is the same as at left (promoter). Pairwise comparisons of other tissues showed similar results (data not shown). (C) Known transcription factor motif analysis of tDMR promoters. We used the JASPAR database (http://jaspar.genereg.net) to search for over-represented transcription factor binding sites in tDMRs. (+) tDMRs that are hypermethylated in the tissue of interest; (−) tDMRs that are hypomethylated in the tissue of interest.











