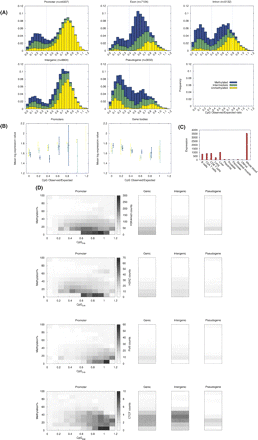
Analysis of somatic DNA methylation profiles. (A) Distribution of data with respect to CpGo/e for the different genome feature categories. The data were operationally categorized into unmethylated (<40%), intermediate (40%–60%), and methylated (>60%). Within the nonpromoter categories, we focused predominantly on CpG islands as annotated in the Ensembl genome browser (NCBI_36). Therefore, the average CpGo/e within the nonpromoter categories in our data set was higher than the genome average (refer to Table 1). However, because probes could not be chosen for all nonpromoter CpG islands, we also randomly selected some CpG-poor nonpromoter regions, and hence, a bimodality of CpGo/e is also observed in some of the nonpromoter categories. Methylation data used in these plots are from whole blood. (B) Comparison of promoter DNA methylation with gene expression across a range of promoter CpGo/e. Whole-blood DNA methylation data (only ROIs overlapping the TSS were used) was correlated with whole-blood genome-wide expression profiles obtained from the GNF SymAtlas database (Su et al. 2004). There were insufficient data for intermediately methylated promoters in the CpGo/e = 1.2 category, and methylated promoters in the CpGo/e ≥ 1 categories. The color code is the same as in A, and error bars represent 95% confidence intervals. (C) Gene expression levels for ICAM3 were obtained from a public database (Su et al. 2004). Expression values represent average difference values computed by Affymetrix software. These values are proportional to mRNA content in the sample. (D) Correlation of DNA methlyation with H3K4me3, H2A.Z, RNA PolII, and CTCF enrichment. DNA methylation data (500 bp ROIs) from our study were correlated with genome-wide enrichment profiles for 20 histone lysine and arginine methylations, H2A.Z, RNA PolII, and CTCF generated by Barski et al. (2007) using Illumina 1G sequencing technology. The remaining 19 comparisons are presented in the Supplementary section. The X-axes represent CpGo/e (there were insufficient data to stratify by CpGo/e in the nonpromoter categories), the Y-axes DNA methylation levels, and the grayscale represents the average tag count for the histone modification or protein indicated. The exon and intron categories were combined into a single “genic” category. Hatched regions indicate that insufficient data were available.











