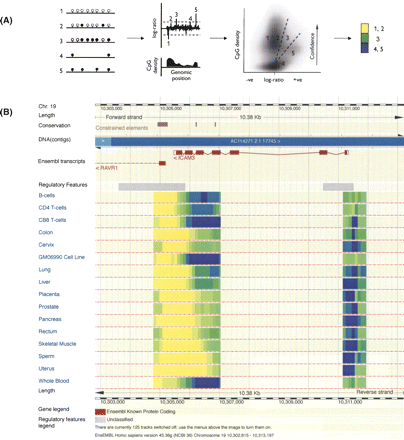
(A) Schematic description of Batman. The left panel shows five hypothetical genomic regions of varying CpG densities and number of methylated CpG sites (filled and empty circles represent methylated and unmethylated CpG sites, respectively). As MeDIP enrichment is proportional to the number of methylated CpG sites, the normalized enrichment ratios of these five hypothetical regions, shown in the second panel, will not accurately reflect the absolute methylation levels at the genomic region of interest (ROI). Batman is based on the observation that the log-ratio MeDIP signal of methylated DNA scales linearly with the number of methylated CpG sites in a sequence. We use a Bayesian deconvolution strategy, taking into account the estimated distribution of DNA fragment lengths, to find the most likely configurations of methylated and unmethylated CpGs in a sequence that explains the observed MeDIP signals. This allows estimation of absolute methylation levels at the ROI. Yellow, green, and blue represent unmethylated, intermediately methylated, and methylated regions, respectively. Batman is described in detail in Down et al. (2008). (B) Integration of Batman-called methylation values into the Ensembl genome browser—screenshot of the data integrated into the Ensembl genome browser (www.ensembl.org). The web display uses a color gradient to show the Batman methylation score for each of the probes in the ROI. The color represents the value of the probe on a sliding scale from 20 (bright yellow) to 80 (dark blue). Probes with Batman values of less than 20 or greater than 80 are colored with the maximum and minimum shades to increase the contrast in the display. Each tissue-type is configured as a dedicated DAS source, allowing the user to select any possible subset of tissues for viewing. Users clicking on a probe will see a small pop-up window, which displays the exact chromosome position of the probe and the Batman score.











