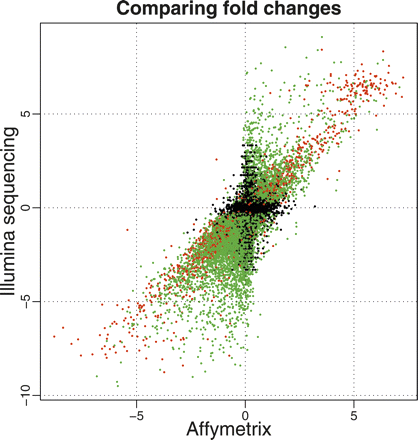
Comparison of estimated log2 fold changes (liver/kidney) from Illumina (Y-axis) and Affymetrix (X-axis). We consider only genes that were interrogated using both platforms and genes where the mean number of counts across lanes was greater than 0 for both the liver and kidney samples. (Red and green dots) Genes called as differentially expressed based on the Illumina sequencing data at an FDR of 0.1%, with a mean number of counts greater than (red) or less than (green) 250 reads in both tissues. (Black dots) Genes not called as differentially expressed based on the Illumina sequencing data. The set of differentially expressed genes that show the strongest correlation between the two technologies seems to be those that are mapped to by many reads (red), while the correlation is weaker for differentially expressed genes mapped to by fewer reads (green).











