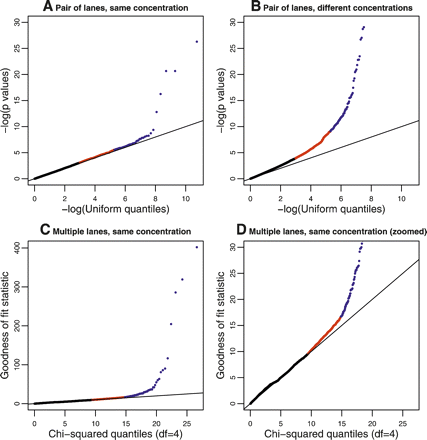
Plots to assess lane effects. Each panel shows a qq-plot comparing the distribution of a statistic (Y-axis) against its theoretical distribution in the absence of a lane effect (X-axis). Deviations from the line y = x indicate the presence of a lane effect. (Points in red) Those above the 95th percentile; (points in blue) those above the 99.5th percentile. (A) A typical result when using P-values derived from a hypergeometric test statistic to compare two lanes used to sequence the same sample at the same concentration. (In this panel, data generated when the kidney sample was sequenced in Run 1, lane 1 and Run 2, lane 2 were used; see Supplemental Fig. 4 for all pairwise comparisons.) (B) Analogous results when comparing two lanes used to sequence the same sample at different concentrations. (In this panel, data generated when the kidney sample was sequenced in Run 1, lane 1 and Run 2, lane 4 were used; see Supplemental Fig. 5 for all pairwise comparisons.) (C,D) Results (on two different scales) when the goodness-of-fit statistic is used to assess the fit of the Poisson model to the kidney data sequenced at a concentration of 3 pM. The liver sample showed a similar pattern (Supplemental Fig. 6).











