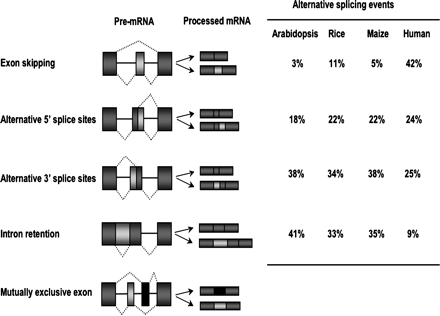
Common types of alternative splice events. The total numbers of EST/cDNAs with validated alignments (similarity ≥ 97% and EST/cDNA coverage ≥ 75%) included in this analysis are: Arabidopsis, 541,594; Rice, 903,022; Maize, 914,822. MAGI3.1 genome data (Fu et al. 2005) was used for maize AS analysis, Transcript isoform determination for all three plant species was carried out with the PASA spliced alignment software described by Haas et al. (2003). The data for human AS event types was reported by Kim et al. (2007)











