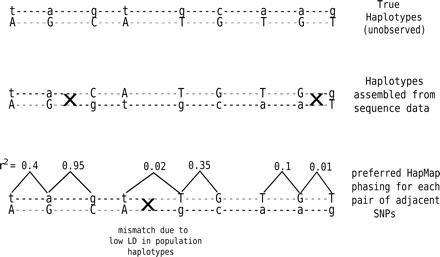
Comparison of haplotypes assembled using sequence data with the preferred HapMap phasing for each pair of adjacent SNPs inferred from the HapMap haplotypes. For three pairs of adjacent SNPs, the phase of the sequence-based haplotypes mismatches the preferred HapMap phasing (indicated by crosses). The first pair shows strong linkage disequilibrium (r2 = 0.95), and therefore, the mismatch is more likely to represent a switch error in the sequence-based haplotypes. For the second pair of SNPs, the sequence-based haplotypes are correct and the mismatch is due to low LD between the SNP pair. For the third pair, LD is again low and the mismatch is due to a switch error in the sequence-based haplotypes.











