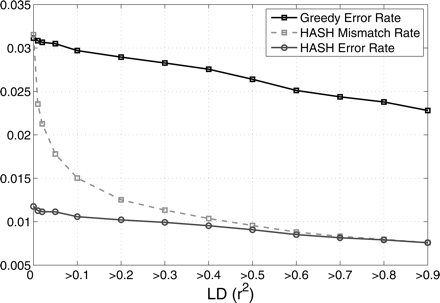
Figure 8.
Mismatch rate and the “adjusted mismatch rate” (error rate) of the HuRef haplotypes estimated by comparison with the CEU HapMap haplotypes. The error rate is plotted as a function of r2, i.e., computed for all pairs of adjacent SNPs with r2 greater than a certain value.











