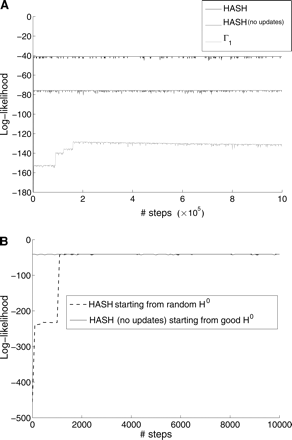
Results of running the MCMC algorithm with different Γ on a fragment matrix with n = 200 columns (from chromosome 22 of HuRef genome). (A) A comparison of the HASH algorithm against two other MCMC algorithms: (1) ℳ(Γ1) and (2) ℳ(Γ) where Γ was computed using the recursive graph-partitioning algorithm G(X). All algorithms were initialized with a random haplotype pair. (B) Comparison of HASH algorithm initialized with a random haplotype against ℳ(Γ) (graph-partitioning) initialized with a good haplotype. Note that we are zooming in on the first 10,000 steps in the iteration.











