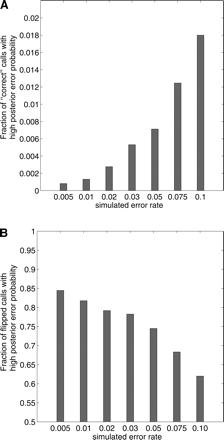
Figure 5.
Fraction of variant calls with a posterior error probability of ≥0.5 using the HASH algorithm for different values of ε. (A) False-positive rate, given by the fraction of “correct” variant calls with high posterior error probabilities. (B) True-positive rate, given as the fraction of “flipped” variant calls with high posterior error-probability.











