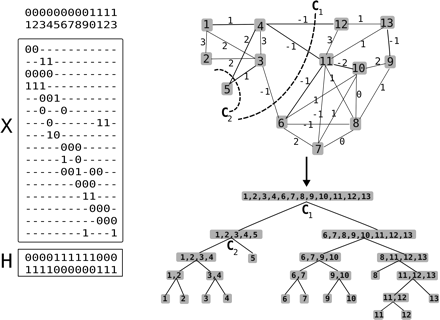
Illustration of the recursive graph-partitioning algorithm for computing Γ. The weighted graph G(X) derived from the fragment matrix X and a haplotype pair H is shown on the top right. The tree structure below demonstrates the recursive partitioning of the columns of X using min-cut computations in the graph G(X). The first cut (C1), partitions the columns of X into two subsets: S = {1, 2, 3, 4, 5} and S̅ = {6, 7, 8, 9, 10, 11, 12, 13}. The second cut (labeled C2), further partitions the subset S into two smaller subsets: {1, 2, 3, 4} and {5}. Γ is obtained from the subsets labeling the nodes of the tree (except the root node).











