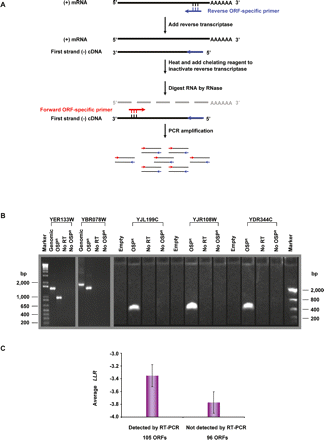
Two-step strand-specific RT-PCR. (A) Schematic diagram of the strand-specific RT-PCR procedure. (B) Electrophoretic analysis of strand-specific RT-PCR products. Reverse ORF-specific primers (OSPR), with sequences complementary to the ORF-coding strand, were used for first-strand cDNA synthesis. Second-step PCR amplifications used a pair of forward (OSPF) and reverse ORF-specific primers (OSPR). As controls, the first step of RT-PCR was performed without reverse transcriptase for detecting contamination by genomic DNA, or without the OSPR primer for detecting residual reverse transcriptase activity in second-step PCR reactions. Two intron-containing verified ORFs, YER133W (genomic DNA length: 1464 bp; coding sequence length: 939 bp) and YBR078W (genomic DNA length: 1737 bp; coding sequence length: 1407 bp), were used to test the strand specificity. An extra control for these two verified ORFs was a standard PCR action using yeast genomic DNA as template and the same pair of ORF-specific primers. The observed difference in the length of PCR products amplified from genomic DNA versus poly(A) mRNA manifested the strand specificity. Strand-specific RT-PCR results of 201 nonoverlapping orphan ORFs were analyzed on 1% agarose E-gel (Invitrogen). Of the reactions 53% (105 ORFs) gave rise to visible RT-PCR products of the expected sizes. Three orphan ORFs, YJL199C (327 bp), YJR108W (372 bp), and YDR344C (444 bp), are shown as examples of successful RT-PCR reactions. (C) Comparison of the average LLR between nonoverlapping ORFs detected and undetected by strand-specific RT-PCR. Error bars, SEM.











