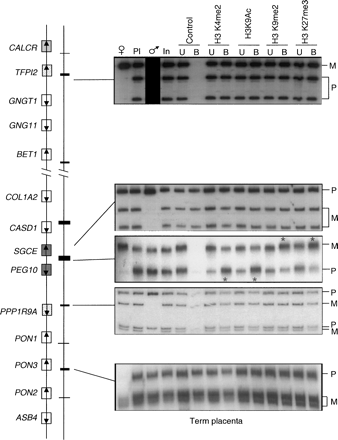
Allele-specific histone methylation and acetylation in human term placenta. DNA extracted from antibody bound (B) and unbound (U) fractions were PCR amplified and followed by single strand confirmation polymorphism (SSCP) to differentiate parental origin. Polymorphism and primer locations are given in Supplemental Table 5. For each bound fraction, we determined the ratio between parental alleles, after correction against allelic ratio of the input chromatin. Asterisks indicate the lanes where, after correlation against the allelic ratio in the input chromatin, the allelic ratios were higher than 2. The only region of allelic enrichment was within the PEG10/SGCE DMR. The paternal relative enrichment for H3K4me2 was 2.3, and 2.1 for H3K9ac. The maternal relative enrichment for H3K9me2 was 3.3; for H3K27me3, 2.7. No allelic enrichment was observed within any other promoter region. Open boxes depict genes that are biallelic in all tissues throughout gestation, whereas dark gray boxes represent the ubiquitously imprinted, paternally expressed genes. Light gray boxes represent genes that are ubiquitously maternally expressed, while hatched boxes are the placental-specific maternally expressed genes. Arrows show the direction of transcription for each gene.











