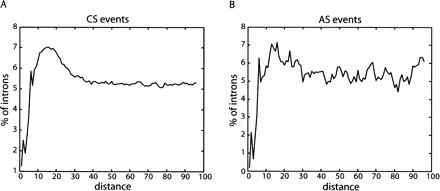
Positional bias of intronic U-rich motifs. Distribution of U-rich (4/5 U’s) motifs in the first 100 nucleotides of intronic sequences downstream of constitutively spliced (CS) (A) and alternatively spliced (AS) (B) exons. U-rich motifs are enriched between nucleotides +6 and +30 of introns downstream of CS and AS exons. The difference in the nature of the plots between CS and AS events (i.e., more smooth for CS events) is due to different data set sizes (109,225 vs. 3872). U-rich motifs are not enriched in the corresponding intronic region downstream of pseudo exons (Supplemental Fig. 1). Other U-rich motifs (3/3, 3/4, 4/4, 5/5, 5/6, 6/6, 6/7, 7/7 Us) were also assessed and showed similar distributions in introns adjacent to constitutive exons, whereas only motifs that have at least one positionally flexible nucleotide showed similar distributions in introns downstream of alternative exons (Supplemental Fig. 1).











