Comparison of several medium- and large-scale motif discovery benchmarks
Click on table to view larger version.
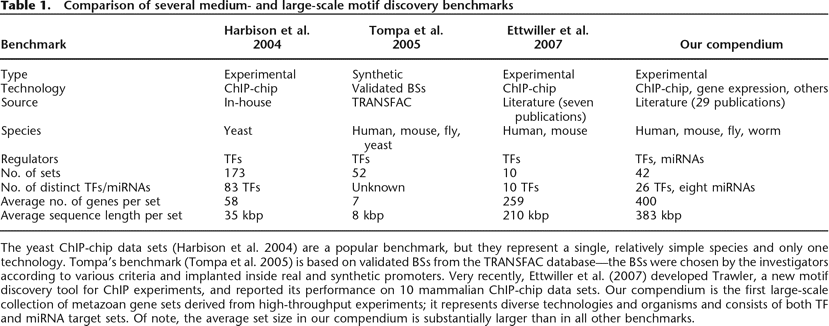
The yeast ChIP-chip data sets (Harbison et al. 2004) are a popular benchmark, but they represent a single, relatively simple species and only one technology. Tompa’s benchmark (Tompa et al. 2005) is based on validated BSs from the TRANSFAC database—the BSs were chosen by the investigators according to various criteria and implanted inside real and synthetic promoters. Very recently, Ettwiller et al. (2007) developed Trawler, a new motif discovery tool for ChIP experiments, and reported its performance on 10 mammalian ChIP-chip data sets. Our compendium is the first large-scale collection of metazoan gene sets derived from high-throughput experiments; it represents diverse technologies and organisms and consists of both TF and miRNA target sets. Of note, the average set size in our compendium is substantially larger than in all other benchmarks.











