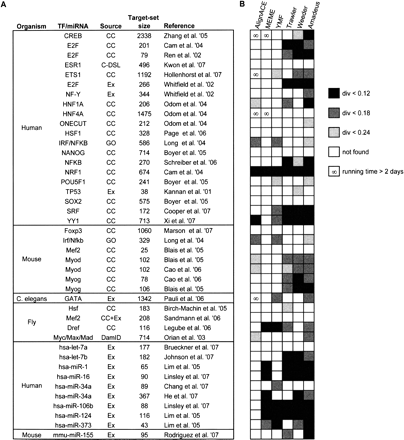
The metazoan target-set compendium and benchmark results on it. (A) The compendium of metazoan TF/miRNA target sets collected from the literature. The “Source” column indicates the experimental procedure or database from which the target set was derived: gene expression microarrays (Ex), ChIP-chip (CC), ChIP-DSL (C-DSL), DamID (van Steensel et al. 2001), or Gene Ontology (GO) database (Ashburner et al. 2000). For additional information and references, see http://acgt.cs.tau.ac.il/amadeus. (B) Performance of motif finding tools on each target set—each successful motif recovery is marked by a gray-shaded box, according to the PWM divergence (darker shades of gray indicate higher similarity of the recovered motif to the one in the literature); the ∞ symbol marks long executions (>48 h) that were aborted. Here, Amadeus was run with the HG enrichment score. The success-rate patterns of the six motif finders are almost identical when comparing different target sets of the same TF. For example, in all three E2F data sets, Amadeus, Weeder, and Trawler are the only tools that recovered the correct motif; in the two Myod sets, Amadeus and Weeder succeeded with PWM divergence cutoff 0.18, AlignACE succeeded with cutoff 0.24, and MEME and YMF failed with all cutoffs. This consistency, observed for all six TFs that are represented by more than one set in our compendium, is not a result of large overlaps between the target sets, as such overlaps were avoided in the construction of the compendium. Instead, it is likely to stem from properties inherent to the TFs, such as the extent and type of their BSs degeneracy.











