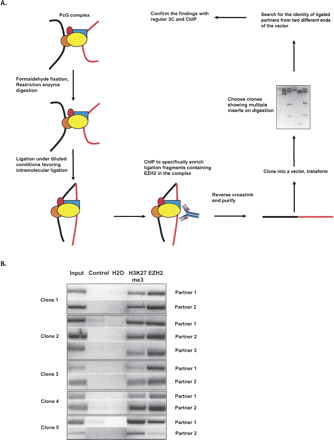
Construction of the 6C assay for EZH2. (A) Outline of the 6C assay. Briefly, cells are subjected to the conventional 3C procedure as described previously (Tolhuis et al. 2002). However, subsequent to the ligation step, the chromatin is subjected to chromatin immunoprecipitation (ChIP) using an antibody against the protein of interest (EZH2, in the present study), followed by ligation of the purified DNA into a cloning vector bearing the sequence overhangs generated in the enzyme digestion used in the 3C assay (EcoRI, in our case) to facilitate insert cloning and further screening. Clones are screened by digestion with the restriction enzyme that was used for the 3C assay, and the ones showing multiple inserts were subjected to sequencing from both directions in the vector to reveal the identity of the partners. (B) EZH2 is bound to the chromatin regions discovered in the 6C assay and is associated with the trimethylation of H3K27 in undifferentiated Tera-2 cells. The ChIP was performed using antibodies against EZH2 and H3K27me3 followed by amplification of the precipitated DNA using primers spanning the EcoRI enzyme sites (from each of the partners) used for 3C assay. The gel shows the results of semiquantitative PCR reactions for each of the partner regions in the five clones under investigation.











