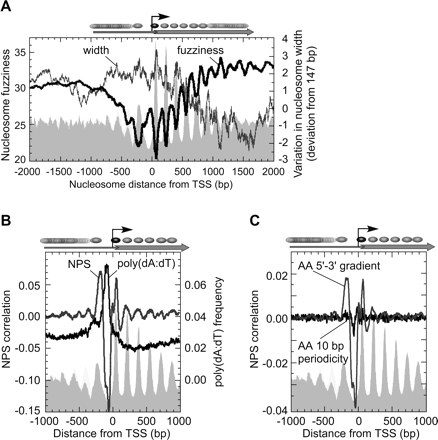
Statistical positioning emanating from barrier nucleosomes. (A) Nucleosome fuzziness and width relative to the TSS. Fuzziness (standard deviation of read locations for those nucleosomes defined by three or more reads) is plotted, against a gray backdrop of Figure 1C. Data are plotted as a moving average of 500 nucleosomes. Deviations from the canonical 147 bp nucleosomal length (in bp) are displayed as a 2000-nucleosome moving average, and represent the distance between the nucleosomal calls made separately on the W and C strand. (B) Averaged NPS correlation score and poly(dA:dT) density for genes aligned by the TSS. The NPS correlation employed an updated AA/TT nucleosomal distribution pattern (Supplemental Fig. S6C). Plot of AAAA/TTTT per bp frequency (11 bp moving average) is shown. Longer poly(dA:dT) tracts gave similar results (data not shown). The gray backdrop reflects nucleosome positions from Figure 1C. NPS scores and fuzziness are not correlated (Supplemental Fig. S7), indicating that the diminishing NPS correlation at distances distal to the −1/+1 nucleosome is unlikely to cause the increased fuzziness shown in panel A. (C) NPS correlation profiles broken out by AA dinucleotide enrichment near 5′ ends vs. 10 bp periodic spacing of AA dinucleotides.











