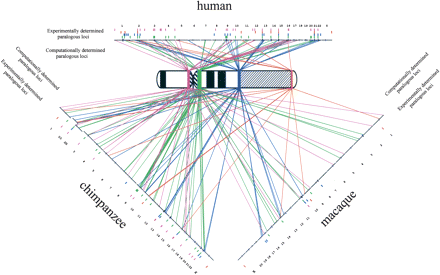
Comparative paralogous duplication pattern of human Y-chromosomal segmental duplications. An ideogram of the G-banded pattern of the normal human Y is shown in the middle. The colored boxes on the chromosome represent the euchromatin/heterochromatin transition regions composed of segmental duplications. The autosomes and the X chromosome are illustrated as small horizontal black lines above (human), rightward diagonally disposed (chimpanzee), and leftward diagonally disposed (macaque). Each chromosome is tagged by the corresponding chromosome number. Heterochromatic regions (constitutive heterochromatin, telomeric caps, and NORs) are indicated as tiny purple boxes on the horizontal lines. All diagonal lines represent pairwise alignments of ≥1 kb and ≥90% nucleotide identity identified by whole-genome sequence comparisons. The small colored boxes in-between the chromosomes and their corresponding numbers display the multi-site pattern from comparative FISH. Concerning the Yq11.1/Yq11.21, Yq11.23/Yq12, and Yq12/PAR2 region, experimental and computational results are 48% consistent for the human, 45% for the chimpanzee, and 30% for the macaque. For the Yp11.1/Yp11.2 region, the concordance decreases to 21% (human), 23% (chimpanzee), and 11% (macaque). Please note that paralogies detected by whole-genome sequence comparisons do not correspond to ancestral duplicon locations.











