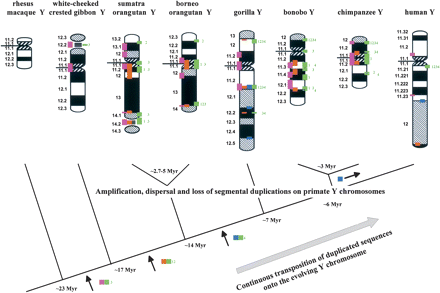
A diagram delineating the evolutionary dynamics of Y-chromosomal segmental duplications. A multi-step process is depicted leading to the current distribution pattern of duplicated sequences on the Y chromosomes of the great and lesser apes. Serial intrachromosomal duplicative transpositions and chromosomal rearrangements generated a mosaic pattern on all great ape Y chromosomes. Continuous interchromosomal transfer of duplicated cassettes provided the basis to develop such a complex structure. The upper row outlines G-banded ideograms for each of the primate Y chromosomes analyzed. Each colored rectangle on a primate Y chromosome indicates the presence of a discrete SD region: (Pink) Yp11.2/Yp11.1; (green) Yq11.1/Yq11.21 (green numbers refer to numbered BAC clones [(1) RP1-85D24, (2) RP11-131M6, (3) RP11-886I11, (4) RP11- 295P22] spanning the human Yq11.1/Yq11.21 transition region); (blue) Yq11.23/Yq12; (orange) Yq12/PAR2. The phylogenetic tree indicates the divergence time in millions of years for each species: ∼6 Mya for the Homo–Pan clade split, ∼3 Mya for chimpanzee–bonobo split, ∼7 Mya for the gorilla, ∼14 Mya for the orangutans; ∼17 Mya for the gibbon, ∼23 Mya for the macaque (Goodman 2005), and ∼2.7–5 Mya for Bornean–Sumatran orangutan split (Steiper 2006). Colored rectangles intermediate to the evolutionary branching points indicate the period of interchromosomal addition or deletion of the respective duplicated sequences.











