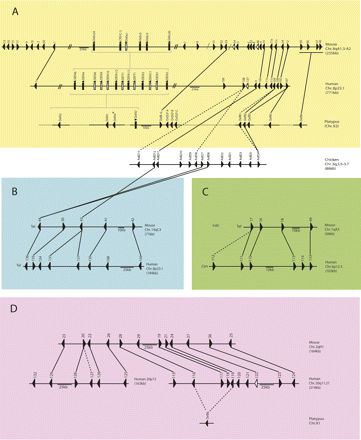
Genomic organization of regions of conserved synteny between alpha- and beta-defensins in eutherians, the platypus, and the chicken, based on previously published data (Belov et al. 2007; Patil et al. 2005). Synteny group A, the most ancient beta-defensin synteny group, contains most of the platypus beta-defensins, with synteny group D, containing DEFB6, arising next. Subsequent duplications and translocations have given rise to synteny groups B and C in the later vertebrate lineage. Eutherian genes with labels containing numbers only are beta-defensins. Chicken beta-defensins are labeled as “AvBD,” and mouse alpha-defensins (cryptidins) are labeled as ‘”Defcr.” Solid lines link orthologs based on the phylogenetic tree (Fig. 2), and broken lines connect paralogs; pseudogenes are indicated by white arrows; slanted lines indicate breaks in the group; slanted dotted lines indicate a mouse beta-defensin cluster that is not mapped on chromosome 8. An asterisk indicates the putative early alpha-defensin. The fragmented nature of the platypus genome sequence (Warren et al. 2008) meant that, although the platypus alpha- and beta-defensins and the OvDLPs were localized to chromosomes using FISH mapping and phylogenetic analysis, contig order could not be determined and is instead inferred based on synteny.











