ALLPATHS assemblies: All edges of size ≥10 kb that contain errors
Click on table to view larger version.
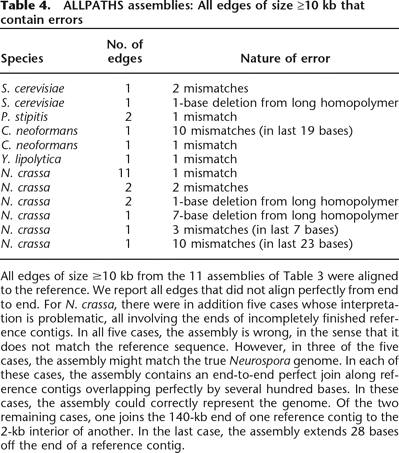
All edges of size ≥10 kb from the 11 assemblies of Table 3 were aligned to the reference. We report all edges that did not align perfectly from end to end. For N. crassa, there were in addition five cases whose interpretation is problematic, all involving the ends of incompletely finished reference contigs. In all five cases, the assembly is wrong, in the sense that it does not match the reference sequence. However, in three of the five cases, the assembly might match the true Neurospora genome. In each of these cases, the assembly contains an end-to-end perfect join along reference contigs overlapping perfectly by several hundred bases. In these cases, the assembly could correctly represent the genome. Of the two remaining cases, one joins the 140-kb end of one reference contig to the 2-kb interior of another. In the last case, the assembly extends 28 bases off the end of a reference contig.











