Number of read pair closures in E. coli using 30-base reads and K = 20
Click on table to view larger version.
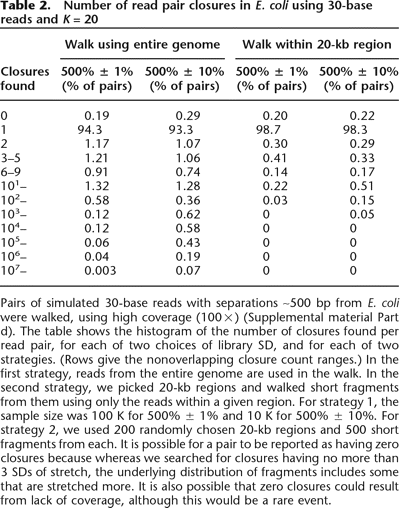
Pairs of simulated 30-base reads with separations ∼500 bp from E. coli were walked, using high coverage (100×) (Supplemental material Part d). The table shows the histogram of the number of closures found per read pair, for each of two choices of library SD, and for each of two strategies. (Rows give the nonoverlapping closure count ranges.) In the first strategy, reads from the entire genome are used in the walk. In the second strategy, we picked 20-kb regions and walked short fragments from them using only the reads within a given region. For strategy 1, the sample size was 100 K for 500% ± 1% and 10 K for 500% ± 10%. For strategy 2, we used 200 randomly chosen 20-kb regions and 500 short fragments from each. It is possible for a pair to be reported as having zero closures because whereas we searched for closures having no more than 3 SDs of stretch, the underlying distribution of fragments includes some that are stretched more. It is also possible that zero closures could result from lack of coverage, although this would be a rare event.











