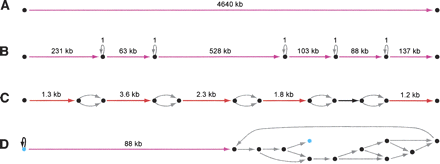
ALLPATHS assemblies of simulated 30-base microreads. Four assembly graphs or parts are shown, with edges color-coded to reflect length: (gray) <100; (black) 100–1000; (red) 1000–10,000; (magenta) >10,000. (Image created using Graphviz [Low 2004].) None of the graph parts have errors, but some have ambiguities. (A) Assembly of E. coli is one edge. (B) 1.1-Mb component from assembly of 21-Mb fungal genome Y. lipolytica. Five ambiguities are seen: All are loops labeled “1” corresponding to mononucleotide runs whose exact length is unknown. (C) Tiny section of component of assembly of diploid human 10-Mb region. Five ambiguities are seen: All are bubbles arising from SNPs, which are intrinsic to the genome. (D) Correct but tangled component of Pichia stipitis assembly. There is a unique path between the two light blue vertices that matches the reference perfectly.











