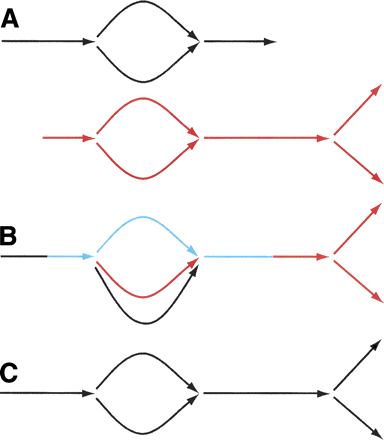
Merger of sequence graphs in ALLPATHS. The process is iterative. It starts with a collection of sequence graphs, and progressively glues them together. Note the simplest case: all the sequence graphs might consist of single edges. Example: (A) two sequence graphs match at graph and sequence level along common portion consisting of bubble extended on both ends; (B) the algorithm identifies a common linear stretch (blue) that extends from a source on one graph to a sink on the other, then glues the graphs along this stretch; however, parallel black and red edges at the bottom are not yet glued; (C) now these edges are zipped up.











