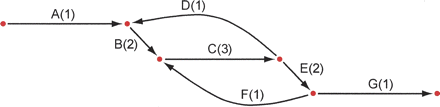
Unipath graph of the 1.8-Mb genome of C. jejuni for K = 6000, which is also the best possible assembly of its genome from unpaired reads of length 6001. The genome was treated as linear to simplify computation. Each unipath is labeled with its number of copies (multiplicity) in the genome and with a letter to facilitate discussion. Formally, the graph also includes a reversed copy corresponding to the reverse-complemented sequence (data not shown). The middle horizontal edge represents a 6.2-kb perfect repeat present three times in the genome. This edge is present exactly because the reads are shorter than the repeat. If the reads were longer than 6.2 kb, then the graph would be a single edge. This graph (along with edge sequences and multiplicities) represents exactly what can be known from the data: There are exactly two ways to traverse the graph from end to end, ABCDBCEFCEG and ABCEFCDBCEG, but it is not possible to know from the data which of these alternatives represents the true genome. When K = 20, one instead has 4161 unipaths in total, and the graph is far too tangled to display, or even to separate the genome from its complemented copy.











